Structural chronobiology
Structural biology of proteins involved in circadian rhythms
Chronobiology has become an increasingly important research area, as the consequences of disturbed biorhythms in humans range from neurophysiological disturbances like sleeplessness and burnout to more severe conditions like metabolic syndrome, diabetes and cancer.
Physiological processes that are regulated on a daily basis, the so-called circadian rhythms, can be found in all kingdoms of life, including humans. Synchronization of these processes to the environmental light-dark cycle requires photosensors, such as the cryptochrome in Drosophila, as well as a strictly regulated periodic inner clock network.
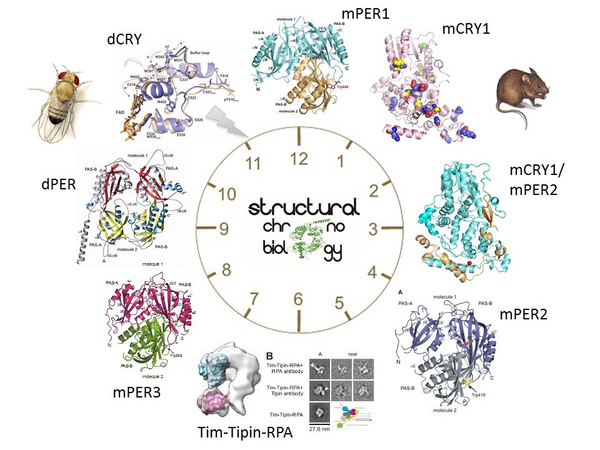
So far, most chronobiological studies have used behavioural, cell biological, and genetic approaches. Structural studies of clock protein functions, and explanations of how maladaptations evolve from disturbances to the periodically oscillating activities and concentrations of clock proteins, are still underrepresented in the field. Hence, our goal is to illuminate the structural bases of clock protein interactions, activities and conformational dynamics underlying these periodic processes.
Our methodical approaches include 3D structure determination by X-ray crystallography as well as biochemical and biophysical studies to analyse protein activities, interactions and conformational changes in solution. Atomic resolution insights into the molecular clockwork and its connection to circadian physiology will pose new biological questions to be addressed by experiments in cell culture and living organisms.
Our structure-function analyses of the circadian photoreceptor Drosophila cryptochrome (dCRY), the clock protein mouse cryptochrome1 (mCRY1) and the complex of mCRY1 with the clock protein PERIOD2 have provided important new insights into the phototransduction mechanism of dCRY as well as into the protein interactions and cellular functions of mammalian cryptochromes (Czarna et al, 2013, Schmalen et al, 2014). Likewise, our crystal structures and structure-based analyses of the PAS (PER-ARNT-SIM) domains of the Drosophila and mammalian PERIOD clock proteins (dPER, mPER1,2,3) have shed light on the versatile functions of these interaction modules in the circadian clock and explain, for example, how a single point mutation in the dPER protein prolongs the fly's day to about 29 hours by affecting protein interactions (Yildiz et al, 2005; Hennig et al, 2009; Kucera et al, 2012). Furthermore, we have determined a Cryo-EM structure of a Timeless (Tim)-Tipin-RPA complex, which plays a role in genome maintenance (Witosch et al, 2014).
