Niehrs Lab
Research Publications Group Members BiographyRegulatory nucleic acid modifications
Our laboratory investigates how regulatory DNA and RNA modifications in vertebrates shape the epigenome and epitranscriptome to control gene expression and define cellular identity during development, differentiation and ageing. Our research spans molecular, cellular and organismal levels, integrating biochemistry, cell biology and bioinformatics. Using Xenopus, murine embryonic stem cells and mice as physiological models, we uncover regulatory mechanisms and assess their phenotypic consequences. This multiscale approach aims to illuminate nucleic acid-based regulation of physiology throughout the lifespan.
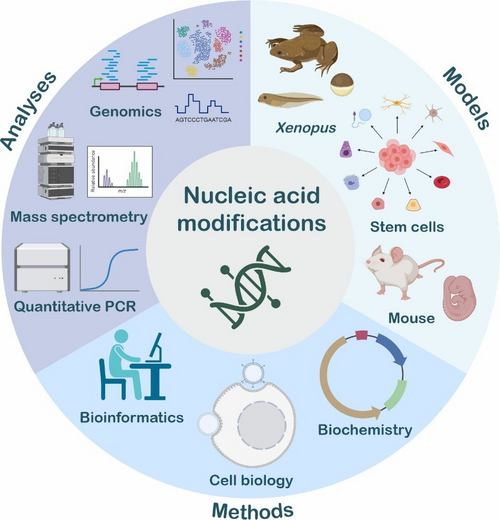
Figure 1: Models, methods and analyses utilised in the Niehrs lab (credit: Johanna M. Schott and Lars Schomacher, created with BioRender.com).
1. Nucleic acid modifications in mammals
Our research in mammals focuses on how DNA modifications shape gene regulation and ageing. We discovered that GADD45a, a non-enzymatic protein, guides active DNA demethylation at specific genomic regions (Barreto et al., Nature, 2007, Schäfer et al., Genes Dev, 2013). In embryonic stem cells, this process can affect stem cell identity (Schüle et al., Genes Dev, 2019), while in mice it protects against premature ageing, directly linking changes in DNA methylation to ageing (Schäfer et al., Genes Dev, 2018). We found that NEIL DNA glycosylases promote oxidative DNA demethylation by accelerating the enzymatic release of the demethylation intermediates 5-formylcytosine and 5-carboxylcytosine (Schomacher et al., Nat Struct Mol Biol, 2016).
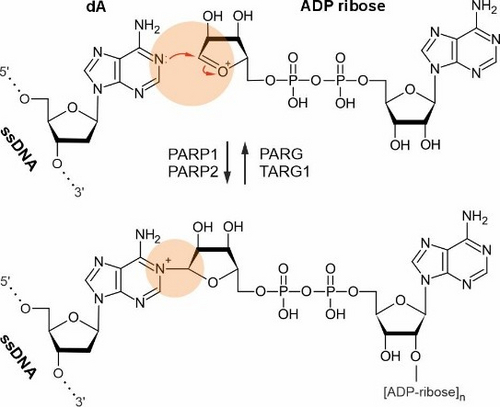
Notably, we established that bases of mammalian DNA can undergo poly-ADP-ribosylation at N1-adenosine residues, an extremely rare but dynamic modification in vivo mediated by PARP1, expanding the known repertoire of DNA base modifications (Musheev et al., Nat Commun, 2022). In addition, we showed that N6-methyl-deoxyadenosine in mammalian DNA is not a classical epigenetic mark; instead, it largely arises from RNA recycling pathways and misincorporation during DNA replication (Musheev et al., Nat Chem Biol, 2020).
2. Nucleic acid modifications in Xenopus development and ageing
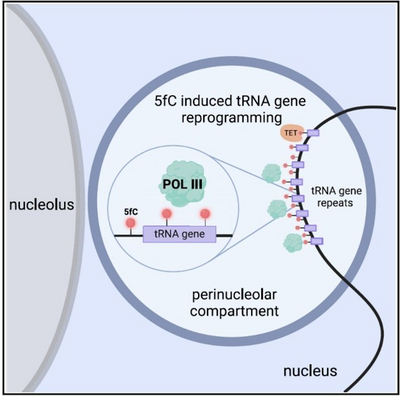
Using Xenopus frogs as a model, we study how DNA modifications regulate early development and ageing. During DNA demethylation, 5-formylcytosine is known as a transient intermediate. We have now discovered that 5-formylcytosine is also an instructive epigenetic mark in its own right. During Xenopus zygotic genome activation, it promotes RNA polymerase III transcription (Parasyraki et al., Cell, 2024). Thus, 5-formylcytosine is only the second instructive epigenetic mark in vertebrates besides 5-methylcytosine.
Also in Xenopus, we created epigenetic clocks by mapping DNA methylation, which predicts biological age, and revealed conserved 5mC ageing signatures are shared between frogs and humans (Zoller et al., Geroscience, 2024). These findings highlight the deep evolutionary conservation of epigenetic regulation and establish frogs as powerful models for understanding both early development and the molecular basis of ageing.
3. Epitranscriptomics and R-loop biology
Complementing our DNA-focused studies, we investigate RNA modifications and R-loop dynamics to understand their roles in gene regulation, epitranscriptome and genome stability, and embryonic development.
To this end, we showed that GADD45a can read R-loops, which are three-stranded structures formed when RNA hybridises to DNA during transcription. R-loops can be regulatory or detrimental to the cell. By binding to regulatory R-loops, GADD45a recruits the DNA demethylation machinery to gene promoters, thereby activating transcription (Arab et al., Mol Cell, 2014, Arab et al., Nat Genet, 2019).
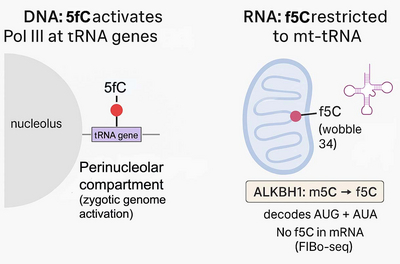
Moreover, we found that across the entire mammalian transcriptome, 5-formylcytosine occurs at only one single position: in mitochondrial tRNAMet. This highlights the biological importance and specificity of rare nucleic acid modifications (Dehnen et al., Nat Commun, 2025). It also demonstrates that despite many similarities between 5-formylcytosine in DNA and RNA, the writer enzymes and locations where it is deposited are radically different between DNA and RNA.
Perspective
Our work reveals how nucleic acid modifications control gene regulation during development and in physiological situations. By linking molecular mechanisms to organismal function, we aim to uncover the fundamental principles of the epigenome and epitranscriptome. Ultimately, we seek to advance a broader understanding of how chemical information embedded in nucleic acids contributes to genome regulation in health and disease.